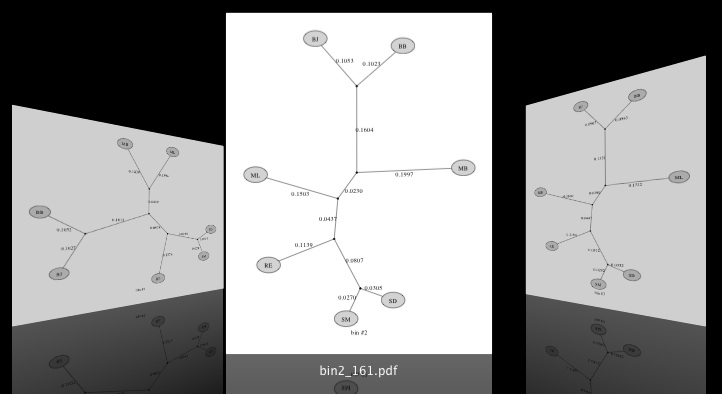
(Above figure) Trees corresponding to the three largest bins resulting from a phybin run. The file binXX_YYY, where XX is the rank of the bin and YYY is the number of trees having that topology.
PhyBin is a simple command line tool that classifies (bins) a set of Newick tree files by their topology. The purpose of it is to take a large set of tree files and browse through the most common tree topologies.

(Above figure) Trees corresponding to the three largest bins resulting from a phybin run. The file binXX_YYY, where XX is the rank of the bin and YYY is the number of trees having that topology.
PhyBin is a command-line program that produces output in the form of text files and pdfs, but to produce pdfs (to visualize trees) the GraphViz program, including the dot command, must be installed on the machine.
The following is a typical invocation of PhyBin:
phybin *.tree -o output_dir/
The input trees can be specified directly on the command-line, or, if the name of a directory is provided instead, all contained files are assumed to be trees in Newick format.
PhyBin, at minimum, produces files of the form output_dir/binXX_YY.tr, one for each bin. If requested, it will also produce visual representations of each bin in the form output_dir/binXX_YY.pdf.
The source code to PhyBin can be downloaded here:
PhyBin is written in Haskell and if you have Haskell Platform installed you can install phybin with this one-liner:
cabal install phybin
PhyBin is also available for download as a statically-linked executable for Mac-OS and Linux:
It should be possible to build it for Windows, but I haven’t done so yet.
In addition to input files and directories, phybin supports a number of command-line options.
As of PhyBin Version 0.1, the -n option is mandatory to specify how many taxa (leaves) are expected in the trees, and input trees with the wrong number of taxa are ignored.
-v or --verboseprint WARNINGS and other information (recommended at first)
-V or --versionshow version number
-o DIR or --output=DIRset directory to contain all output files (default “./”)
-g or --graphbinsuse graphviz to produce .dot and .pdf output files named bin1_N., bin2_M., etc
-d or --drawbinslike -g, but open GUI windows to show a tree for each bin
-w or --viewfor convenience, “view mode” simply displays input Newick files without binning
-n NUM or --numtaxa=NUMexpect NUM taxa for this dataset
-p NUM or --nameprefix=NUMLeaf names in the input Newick trees are usually gene names, not taxa. It is typical to extract taxa names from genes. This option extracts a prefix of NUM characters to serve as the taxa name.
-s STR or --namesep=STRAn alternative to —nameprefix, STR provides a set of delimeter characters, for example ’-’ or ‘0123456789’. The taxa name is then a variable-length prefix of each gene name up to but not including any character in STR.
-m FILE or --namemap=FILEEven once prefixes are extracted it may be necessary to use a lookup table to compute taxa names, e.g. if multiple genes/plasmids map onto one taxa. This option specifies a text file with find/replace entries of the form “
Authors: Irene Garcia Newton and Ryan Newton
Contact email: inewton wellesley edu with “at” and “dot” inserted.