Unified Brain Surface and Volume Registration
Maks Ovsjanikov4 Lilla Zöllei2 John Guttag1 Bruce Fischl2* Adrian V. Dalca1,2*
Abstract
Accurate registration of brain MRI scans is fundamental for cross-subject analysis in neuroscientific studies. This involves aligning both the cortical surface of the brain and the interior volume. Traditional methods treat volumetric and surface-based registration separately, which often leads to inconsistencies that limit downstream analyses. We propose a deep learning framework, NeurAlign, that registers 3D brain MRI images by jointly aligning both cortical and subcortical regions through a unified volume-and-surface-based representation. Our approach leverages an intermediate spherical coordinate space to bridge anatomical surface topology with volumetric anatomy, enabling consistent and anatomically accurate alignment. By integrating spherical registration into the learning, our method ensures geometric coherence between volume and surface domains. In a series of experiments on both in-domain and out-of-domain datasets, our method consistently outperforms both classical and machine learning-based registration methods—improving the Dice score by up to 7 points while maintaining regular deformation fields. Additionally, it is orders of magnitude faster than the standard method for this task, and is simpler to use because it requires no additional inputs beyond an MRI scan. With its superior accuracy, fast inference, and ease of use, NeurAlign sets a new standard for joint cortical and subcortical registration.
Method
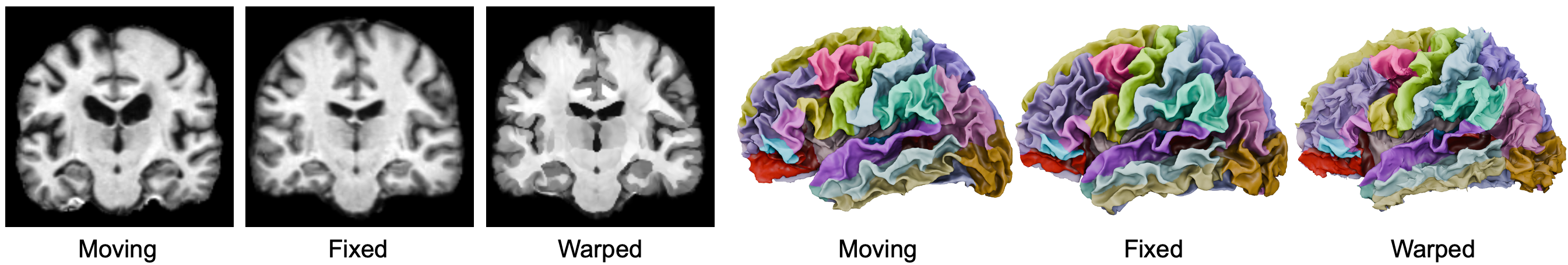
NeurAlign is a model for pairwise registration of subcortical and cortical structures in brain MRI scans. The cortex is a thin, highly folded surface responsible for high-level functions of the brain. Aligning the cortex is ill-suited to standard Euclidean, grid-based registration: homologous folds cannot be reliably identified by image intensity alone, and the cortex lies on a curved 2D manifold that a 3D volumetric grid is poorly equipped to represent.
NeurAlign learns a joint representation of the interior brain volume and cortical surface. The key idea is to use the sphere as an intermediate domain for cortical alignment: we simultaneously train a 3D volumetric registration network and a 2D spherical registration network, producing deformation fields \(\varphi\) and \(\psi\) respectively. The two networks are coupled via a cortical consistency loss \(\mathcal{L}_\text{cons}\) that explicitly encourages the volumetric deformation field \(\varphi\) on the cortical ribbon to agree with the spherical deformation \(\psi\) lifted back to 3D space.
An advantage of our approach is that the spherical and cortical consistency losses are used only during training. At inference time, NeurAlign, requires only a pair of 3D MRI scans—no surface meshes nor segmentation masks are needed. This can save hours of costly processing to extract meshes required by other methods.

NeurAlign jointly trains two networks: a 3D volumetric U-Net that predicts a diffeomorphic deformation field \(\varphi\), and a 2D U-Net that predicts a deformation \(\psi\) on the inflated cortical surface. A cortical consistency loss \(\mathcal{L}_\text{cons}\) couples the two networks during training. At inference time, only the volumetric network is used—no meshes are required.
Results
We train a single model using T1w scans from the OASIS and ADNI datasets. We evaluate our model on held-out test subjects from these datasets as well as on out-of-domain subjects from the MBB and IXI datasets. We measure cortical and subcortical structure overlap using the Dice score, and deformation regularity using the Jacobian determinant. We compare against the most widely-used classical method, CVS, two foundational models: SynthMorph and uniGradICON, and against VoxelMorph, and FireANTs.
NeurAlign Substantially Improves Cortical Alignment Without Sacrificing Subcortical Alignment
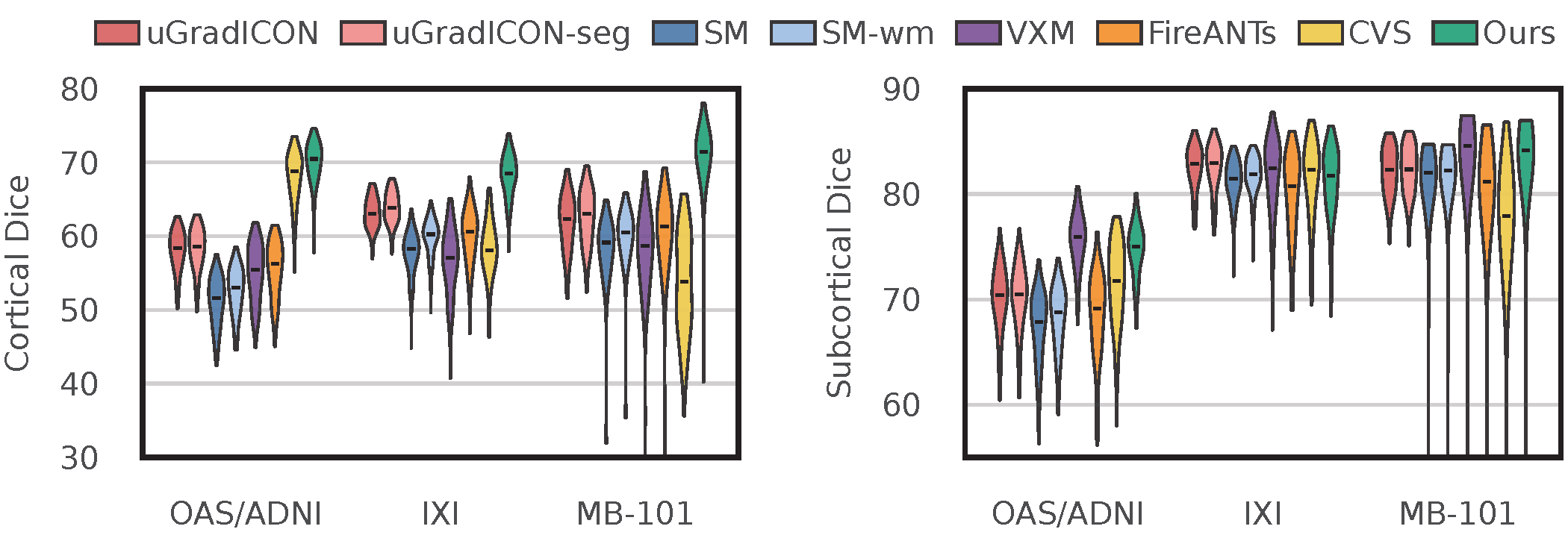
NeurAlign (green) achieves substantially higher cortical Dice across all three datasets, while preserving subcortical performance.
Challenging Cortical Folds are Matched with Smooth, Regular Deformation Fields
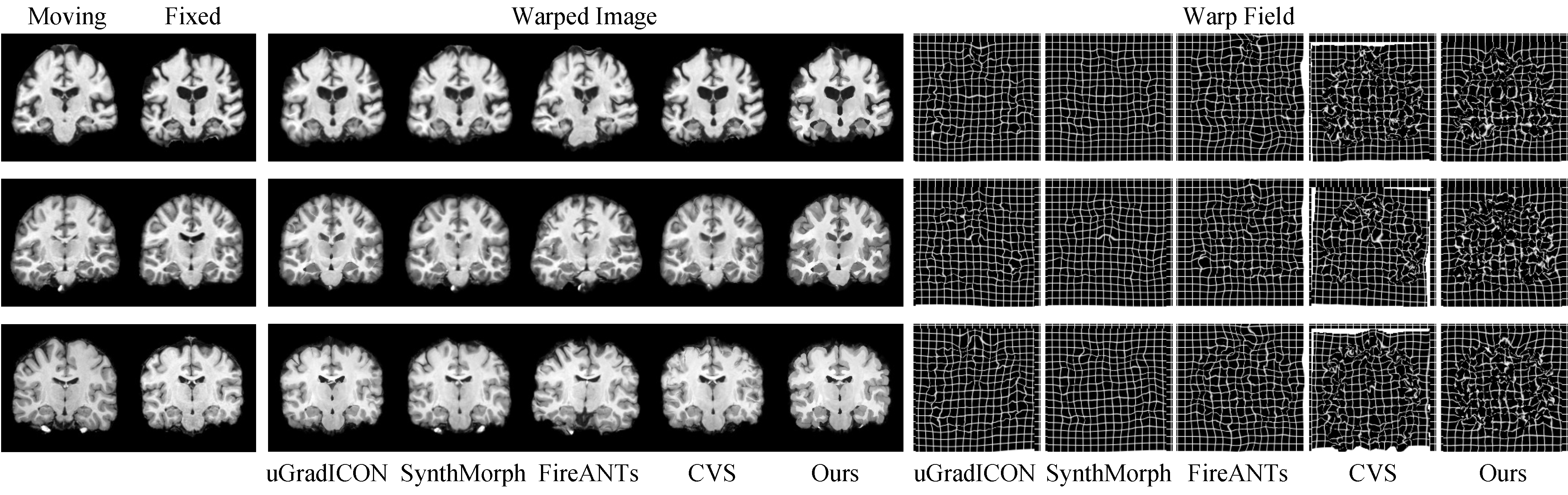
Example registrations for three image pairs. Columns show the moving and fixed images, followed by warped images produced by baselines and our method. Corresponding deformation fields are visualized on the right. Our method yields accurate alignments with smooth and regular warp fields.
Paper
S. Mazdak Abulnaga, Andrew Hoopes, Malte Hoffmann, Robin Magnet, Maks Ovsjanikov, Lilla Zöllei, John Guttag, Bruce Fischl*, Adrian V. Dalca*
Unified Brain Surface and Volume Registration
International Conference on Learning Representations (ICLR), 2026
arXiv | BibTeX
@inproceedings{abulnaga2026neuralign,
title={Unified Brain Surface and Volume Registration},
author={Abulnaga, S Mazdak and Hoopes, Andrew and Hoffmann, Malte and Magnet, Robin and Ovsjanikov, Maks and Z{\"o}llei, Lilla and Guttag, John and Fischl, Bruce and Dalca, Adrian},
booktitle={Proceedings of the International Conference on Learning Representations},
year={2026}
}
