Biography
Hello! I’m a 5th year PhD student in EECS at MIT, co-advised by David Sontag in MIT CSAIL and Gaddy Getz at the Broad Institute. I develop machine learning tools to better understand health and disease. My main focus is on understanding and leveraging RNA-sequencing data, especially to improve precision medicine in cancer. Currently, I’m interested in deep representation learning, generative models, and multimodal integration/translation for biological data.
-
PhD in Electrical Engineering & Computer Science, expected 2025
Massachusetts Institute of Technology
-
MSc in Biomedical Engineering, 2016
Columbia University
-
BA in Physics, 2014
Yeshiva University
Featured Publications
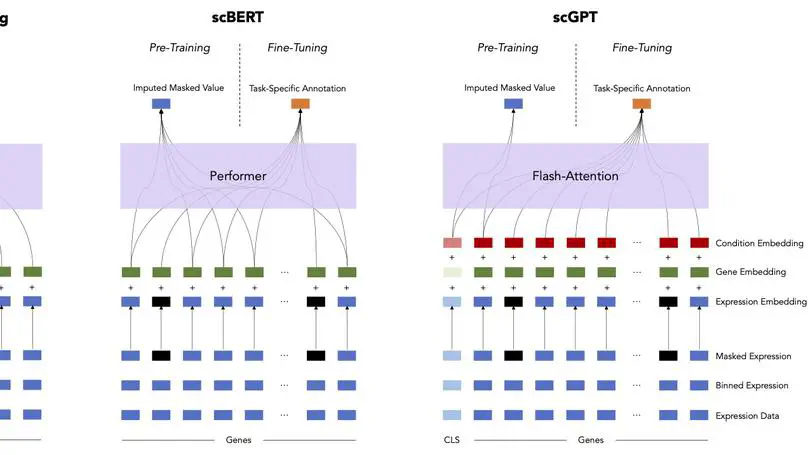
Large-scale foundation models, which are pre-trained on massive, unlabeled datasets and subsequently fine-tuned on specific tasks, have recently achieved unparalleled success on a wide array of applications, including in healthcare and biology. In this paper, we explore two foundation models recently developed for single-cell RNA sequencing data, scBERT and scGPT. Focusing on the fine-tuning task of cell type annotation, we explore the relative performance of pre-trained models compared to a simple baseline, L1-regularized logistic regression, including in the few-shot setting. We perform ablation studies to understand whether pretraining improves model performance and to better understand the difficulty of the pre-training task in scBERT. Finally, using scBERT as an example, we demonstrate the potential sensitivity of fine-tuning to hyperparameter settings and parameter initializations. Taken together, our results highlight the importance of rigorously testing foundation models against well established baselines, establishing challenging fine-tuning tasks on which to benchmark foundation models, and performing deep introspection into the embeddings learned by the model in order to more effectively harness these models to transform single-cell data analysis. Code is available at https://github.com/clinicalml/sc-foundation-eval.
Healthcare providers are increasingly using machine learning to predict patient outcomes to make meaningful interventions. However, despite innovations in this area, deep learning models often struggle to match performance of shallow linear models in predicting these outcomes, making it difficult to leverage such techniques in practice. In this work, motivated by the task of clinical prediction from insurance claims, we present a new technique called Reverse Distillation which pretrains deep models by using high-performing linear models for initialization. We make use of the longitudinal structure of insurance claims datasets to develop Self Attention with Reverse Distillation, or SARD, an architecture that utilizes a combination of contextual embedding, temporal embedding and self-attention mechanisms and most critically is trained via reverse distillation. SARD outperforms state-of-the-art methods on multiple clinical prediction outcomes, with ablation studies revealing that reverse distillation is a primary driver of these improvements. Code is available at https://github.com/clinicalml/omop-learn.
Contact
- rboiar [at] mit [dot] edu